|
4/11/2023 0 Comments Sqlite browser nt authority 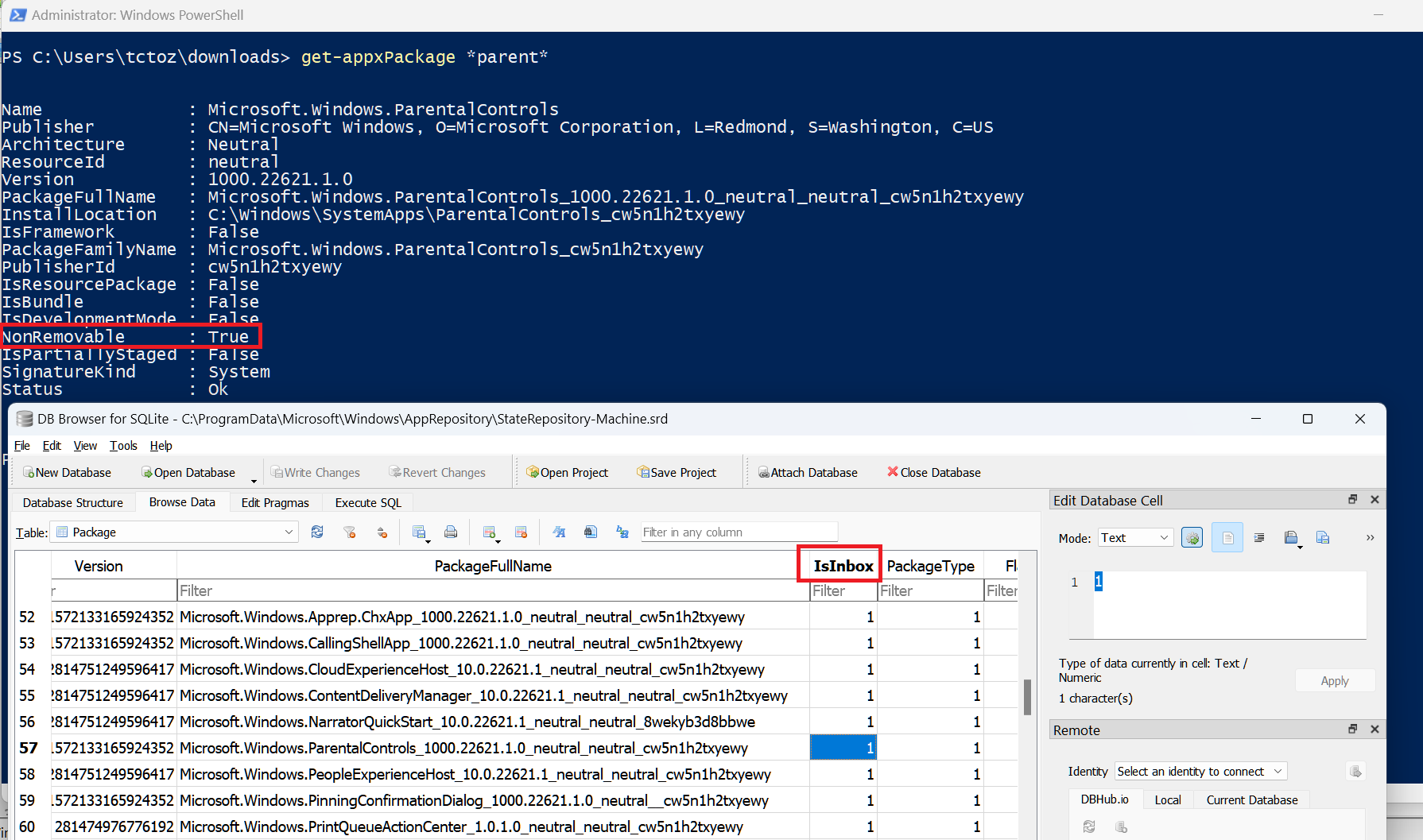
Thus, a bacterial genome, ~4M characters, with 4 sequences takes about 14 seconds to process entirely in 4 CPUs. Multiple sequences are accepted as input, each sequence being analyzed in one CPU. For each input genome, a database SQLite is generated holding the absolute frequencies of all existing strings in the genome up to a user defined length, and their reverse complements. The development was carried out in Python, using packets for handling sequences in FASTA format and the required calculations. In this work, we circumvent this issue, implementing our statistical test in an efficient and multi-processing computer program. One obstacle to test all available genomes with this statistical test was the efficiency of the computational implementation. In a previous work, we have developed a statistical hypothesis test for the generalized second Chargaff parity rule for any particular string in a genome. the frequency of its reverse complement in the same strand. The generalization states that the frequency of a string of a particular length is similar to. Although the validity of the second rule is still in debate and the biological cause is unknown, a generalized form of the second parity rule has already been proposed. an integrated system for constructing Web applications that supports the creation of Web pages by non - programmers WebWriter includes a direct manipulation Web page editor, the WebWriter Editor, which runs in a Web browser as a CGI service, and the WebWriter Page Generator, which creates new pages as an application runs As in HyperCard, users create a Web application as a stack of pages, where each page can contain output regions that are filled in at runtime by a script This paper describes the WebWriter system, issues of server - based authoring tools, and some example applications Read moreĬhargaff’s second parity rule holds that for each of the two DNA strands in a genome, the %A is similar to %T and %G is similar to %C. 
59 | I S B View full-textĪbstract Constructing server - based Web applications requires creating both Web pages and programs that generate Web pages This requires a knowledge of the Hypertext Markup Language (HTML), the Common Gateway Interface (CGI) protocol, and a programming language, such as C++, Python, or Perl While this is not a barrier for programmers, it is for non - programmers This paper describes WebWriter. The tool can be easily integrated into PBS and SLURM scripts that are provided in the package, enabling users to run multiple large sized BLAST+ outputs in high performance computing facilities. This allows a user defined extraction of gene and protein sequences in FASTA format followed by downstream analysis using graphical representations, all without the need for complex programming skills or licensed software. The package primarily retrieves query hits along with their predicted protein or gene id's. We present a python based command line tool A.L.B.I which can be used to parse and analyse large outputs from standalone BLAST+ runs. A prevalent way to discover meaningful relationship from such data is primarily achieved through BLAST similarity. Additionally, the availability of high performance computing facilities has made it well within the reach of biologists to generate and analyse gigabytes of NGS data such as metagenomic reads, within hours. Low cost next generation sequencing is leading us into a "big data" era. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
